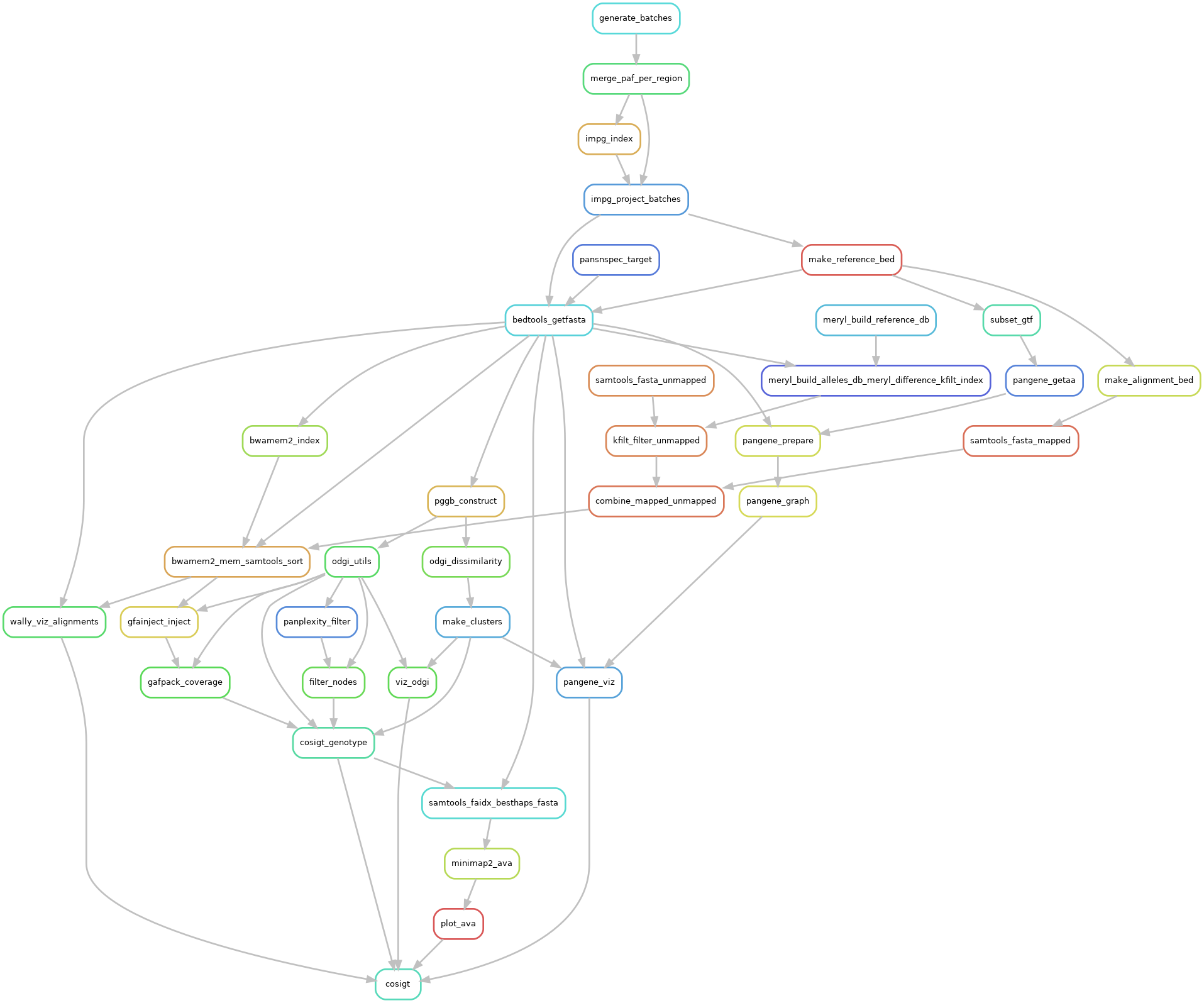
# Workflow
The rulegraph (opens new window) below is a compact representation of cosigt workflow (cosigt rule, master branch)
This section outlines the core steps and logic behind the cosigt pipeline. For each region of interest, the workflow assigns each sequenced sample a pair of haplotypes selected from a given set of assemblies through the following steps:
- Assembly alignment and coordinate projection. Assembled haplotypes are aligned to the reference genome, and target regions are lifted over to identify homologous loci across all haplotypes
- Local pangenome graph construction. Sequences corresponding to target regions are extracted and assembled into a local pangenome graph that captures the haplotype differences
- Graph-based feature extraction. The pipeline computes structural similarity clustering among haplotypes and generates path-specific node coverage vectors representing each haplotype's traversal pattern through the graph
- Read mapping and coverage profiling. Short reads from sequenced samples are mapped to assembly blocks and projected onto the graph to generate sample-specific node coverage vectors
- Cosine similarity-based genotyping. Sample coverage vectors are compared against all possible haplotype pair combinations using cosine similarity, with the highest-scoring pair assigned as the genotype
# Haplotypes alignment
The pipeline begins by aligning the assembled haplotypes (queries) to their corresponding reference genome (target) using minimap2 (opens new window). Minimap2 is executed with the asm20 preset, which is optimized for assembly-to-reference alignment and permits continuous alignment across segments when sequence divergence does not exceed 20%, even within complex regions.
# Target region liftover
Following the initial alignment, the workflow leverages an implicit pangenome graph model - implemented in the impg repository (opens new window) - to lift over coordinates of regions of interest from the target sequence to the queries. This step identifies homologous loci across all haplotypes that map to a specified target region. The alignment coordinates are subsequently filtered and extended to ensure comprehensive coverage of the region of interest.
# Graph construction
With the coordinates of the target regions established, the pipeline extracts the corresponding sequences from both the target and the queries, then uses the pangenome graph builder pggb (opens new window) to construct a local pangenome graph. In previous work (opens new window), collapsing regions with copy-number events into single regions of the graph proved beneficial. Therefore, the pipeline executes pggb with the -c parameter set to 2 by default, enabling construction of a graph that accurately represents variation in loci with high copy number variation.
# Structural clustering
The pipeline implements a strategy to identify structural clusters in the extracted haplotypes. Using the pggb graph, a jaccard-based (opens new window) distance matrix is computed between all haplotype pairs using odgi (opens new window) similarity (scores range from 0 to 1, where 0 indicates identical node traversal patterns and 1 indicates completely disjoint paths). A clustering strategy based on DBSCAN (opens new window) is then applied, with the minPoints parameter set to 1, and the epsparameter dynamically computed. This approach groups haplotypes by structural similarity, providing insights into the major structural patterns present in the dataset.
# Path-specific node coverage
Our workflow uses odgi (opens new window) paths to compute the coverage of each path over the nodes in the graph:
- If a path crosses a certain node just once, its coverage over that node will be
1 - If a path does not cross a certain node, its coverage over that node will be
0 - If a path loops multiple times over a certain node, its coverage over that node will be
>=2
The path coverage information is crucial for the subsequent haplotype deconvolution step.
# Reads-to-assembly alignment
For each sample, short reads spanning regions of interest are extracted from the initial alignment to the linear reference and mapped against the corresponding assembly blocks. Additionally, reads that were originally unmapped are extracted from the initial alignment and filtered through kfilt (opens new window), retaining only those containing at least a 31-mer matching the region of interest. These reads are also mapped against the corresponding assembly blocks.
Mapping is performed using the bwa-mem2 (opens new window) mem algorithm for modern genomes. For ancient genomes, we use bwa (opens new window) aln instead, with parameters optimized for ancient DNA as suggested in this publication (opens new window) (bwa aln -l 1024 -n 0.01 -o 2). The pipeline retains alternative alignment hits to ensure comprehensive coverage of repetitive regions.
# Sample-specific node coverage
The alignments generated in the previous step are projected onto the graph using gfainject (opens new window). Read coverage over the graph nodes is then calculated using gafpack (opens new window). The pipeline incorporates the --len-scale parameter to normalize coverage based on node length and the --weight-queries parameter to account for multiple alignments of the same read, ensuring accurate representation of coverage in repetitive regions.
# Haplotype deconvolution
In the final step of the pipeline, the coverage vector of each sample is compared to a set of pre-computed haplotype coverage vectors. These haplotype vectors are generated by pairing all path vectors over the graph. The comparison is performed using cosine similarity (opens new window), which measures the similarity between two vectors as their dot product divided by the product of their lengths. The haplotype pair with the highest similarity to the short-read vector is assigned as the genotype for each sample in each genomic region. For haploid regions, the same principle applies, but the sample coverage vector is compared to each individual path coverage vector rather than pairs.
# Parallelization
The cosigt pipeline is designed to efficiently utilize available computational resources through extensive parallelization:
- The Assembly-to-Reference Alignment runs in parallel for each chromosome and each assembly sample. The pipeline requires assembled contigs to follow the PanSN (opens new window) specification (
sample#haplotype#contig). All haplotypes sharing the same assembly sample identifier are aligned to their corresponding reference chromosome in a single process. For example, with contigs from 10 different assembly samples and regions from 2 different chromosomes, the workflow can spawn up to 20 concurrent processes during this alignment step. - The Target Region Liftover and the different Graph Analysis operations run in parallel by region, including:
- The Sequencing Sample Processing stages of the pipeline are parallelized by both sequencing sample and region:
- Short-read alignment to assembled contigs (with the exception of unmapped read extraction, which is performed at the sample level)
- Projection of short-read alignments into graph space
- Haplotype deconvolution and genotyping of sequencing samples
This multi-level parallelization strategy enables efficient processing of large cohorts with complex genomic regions, maximizing throughput on high-performance computing infrastructure.