# Use Cases
This section provides a comprehensive end-to-end example of how to run the cosigt (opens new window) pipeline. The rationale of the pipeline is explained in greater detail in the → workflow section.
# Data Acquisition
We create a minimal test dataset to demonstrate cosigt functionality using publicly available data. Let's start by creating a test directory:
mkdir cosigt_test
cd cosigt_test
# Required Tools
Several tools are necessary to complete the following sections. The most straightforward approach is to install them using conda (opens new window) or similar package managers, as shown below.
micromamba create \
-p use_cases_env \
-c bioconda \
-c conda-forge \
samtools \
odgi \
htslib \
agc \
mash \
bedtools \
impg
micromamba activate $PWD/use_cases_env
# Sequencing Reads
The cosigt pipeline requires sequencing reads previously aligned to a reference genome in standard bam (opens new window) or cram (opens new window) format. We can obtain suitable alignment files from the 1000 Genomes Project (opens new window):
mkdir alignments
cd alignments
# Download the complete sample index
wget https://ftp-trace.ncbi.nlm.nih.gov/1000genomes/ftp/1000G_2504_high_coverage/additional_698_related/1000G_698_related_high_coverage.sequence.index
# Select a couple of samples
grep -E "HG00438|HG01952" 1000G_698_related_high_coverage.sequence.index | cut -f 1 > 1000G.selected.txt
# Add their index files
sed 's/$/.crai/' 1000G.selected.txt >> 1000G.selected.txt
# Download the selected files
wget -i 1000G.selected.txt
cd ..
# Reference Genome
A reference genome is required for two purposes: extracting region-specific reads from the original alignments (if in cram format) and serving as one of the assemblies against which samples are compared. In this example, reads are aligned to GRCh38 (opens new window):
mkdir reference
cd reference
# Download the reference genome
wget ftp://ftp.1000genomes.ebi.ac.uk/vol1/ftp/technical/reference/GRCh38_reference_genome/GRCh38_full_analysis_set_plus_decoy_hla.fa
# Download its index
wget ftp://ftp.1000genomes.ebi.ac.uk/vol1/ftp/technical/reference/GRCh38_reference_genome/GRCh38_full_analysis_set_plus_decoy_hla.fa.fai
cd ..
# Genome Assemblies
The cosigt pipeline begins with individual haplotypes to construct pangenome graphs of regions targeted for genotyping. Our workflow expects haplotypes to be grouped by their corresponding reference chromosome. Below are two alternative strategies for generating such chromosome-specific assemblies, which users can choose based on their available data.
# Option 1: Starting from a Chromosome-Specific Graph
One approach is to extract haplotypes for specific chromosomes from existing chromosome-specific pangenome graphs. For example, Year 1 (opens new window) of the Human Pangenome Reference Consortium (HPRC) (opens new window) provides such data (opens new window). We'll limit our analysis to chr4 and chr6 since these contain our regions of interest:
mkdir assemblies_a
cd assemblies_a
# Download chromosome-specific pangenome graphs
wget https://s3-us-west-2.amazonaws.com/human-pangenomics/pangenomes/freeze/freeze1/pggb/chroms/chr4.hprc-v1.0-pggb.gfa.gz
wget https://s3-us-west-2.amazonaws.com/human-pangenomics/pangenomes/freeze/freeze1/pggb/chroms/chr6.hprc-v1.0-pggb.gfa.gz
# Download AGC (Assembly Graph Container)
wget https://zenodo.org/record/5826274/files/HPRC-yr1.agc?download=1 -O HPRC-yr1.agc
# Process each chromosome to subset HG00438, HG01952, plus 8 additional samples
for chr in chr4 chr6; do
# Extract all contig IDs
zgrep "^P" "$chr.hprc-v1.0-pggb.gfa.gz" | cut -f2 | sort -u > "${chr}.all_ids.txt"
# Subset to our samples of interest
grep -E "^HG00438#|^HG01952#" "${chr}.all_ids.txt" > "${chr}.subset.txt"
# Add 8 additional samples for diversity
grep -v -E "^HG00438#|^HG01952#" "${chr}.all_ids.txt" \
| cut -d'#' -f1 \
| sort -u \
| head -n 8 \
| while read id; do grep "^$id#" "${chr}.all_ids.txt"; done >> "${chr}.subset.txt"
rm "${chr}.all_ids.txt"
# Extract FASTA sequences
cat "${chr}.subset.txt" | while read f; do
agc getctg HPRC-yr1.agc $f | bgzip -@ 8 >> "${chr}.subset.fasta.gz"
done
rm "${chr}.subset.txt"
# Index the FASTA file
samtools faidx "${chr}.subset.fasta.gz"
done
cd ..
TIP
This represents just one possible strategy for subsetting assemblies from chromosome-specific pangenome graphs. Alternatively, you can subset the initial .gfa to paths of interest using odgi paths -i GRAPH.GFA -K PATHS.TXT -o SUBGRAPH.OG and then extract the assemblies as FASTA from the subgraph with odgi paths -i SUBGRAPH.OG -f > SUBSET.FASTA.
# Option 2: Starting from Individual Haplotypes
An alternative approach is to determine which reference chromosome each sample-specific haplotype most likely belongs to using Mash distances (opens new window). This method doesn't require chromosome-specific graphs and can start directly from individual assemblies:
mkdir assemblies_b
cd assemblies_b
# Reference to AGC from previous step
agc="../assemblies_a/HPRC-yr1.agc"
# Extract individual IDs
ids=$(cut -d "#" -f 1,2 ../assemblies_a/*fai | sed 's/#/./g' | sort | uniq)
# Download corresponding FASTA files
for id in $ids; do
agc getset $agc $id | bgzip -@ 8 > $id.fasta.gz
samtools faidx $id.fasta.gz
done
# Create a sketch of the reference
mash sketch -s 100000 -p 8 -i ../reference/GRCh38_full_analysis_set_plus_decoy_hla.fa
for id in $ids; do
# Calculate distances (each contig vs reference chromosomes)
mash dist ../reference/GRCh38_full_analysis_set_plus_decoy_hla.fa.msh "$id.fasta.gz" -i -s 100000 -p 8 > "$id.tsv"
# Identify contigs with chr4/chr6 as best matches
cut -f 2 "$id.tsv" | sort -u | while read f; do
best=$(grep -w "$f" "$id.tsv" | sort -k3n | head -1)
chr=$(echo "$best" | awk '{print $1}')
hap=$(echo "$best" | awk '{print $2}')
if [[ "$chr" == "chr4" || "$chr" == "chr6" ]]; then
samtools faidx "$id.fasta.gz" "$hap" | bgzip -@ 8 >> "${chr}.subset.fasta.gz"
fi
done
rm "$id.tsv"
done
# Index the resulting chromosome-specific files
for chr in chr4 chr6; do
samtools faidx "${chr}.subset.fasta.gz"
done
# Regions of Interest
One final piece of information required for the pipeline is a list of target regions located on the chromosomes for which we have assemblies (chromosomes 6 and 4 in this example). Notable examples include the C4 (opens new window) and GYPA (opens new window) loci:
mkdir regions
cd regions
# GYPA region
echo -e "chr4\t144109303\t144140751\tGYPA" > roi.bed
# C4 region
echo -e "chr6\t31972057\t32055418\tC4" >> roi.bed
cd ..
These coordinates serve dual purposes: subsetting the initial assemblies and extracting relevant reads from alignment files.
# Incorporating alternative contigs
When working with references that include alternative contigs (such as the 1000 Genomes alignment reference from the previous section), one should extend the initial BED file to capture reads mapped to these alternate sequences. While not essential for regions like those above, this can be critical for loci such as HLA genes on chr6, which have numerous alternative representations in the reference assembly. The snippet below illustrates how to create such a BED:
cd regions
echo -e "chr6\t29942253\t29945755\tHLA-A
chr6\t31354119\t31357442\tHLA-B
chr6\t31268997\t31272351\tHLA-C
chr6\t32636716\t32643200\tHLA-DQA1
chr6\t32660030\t32667132\tHLA-DQB1
chr6\t32578768\t32590341\tHLA-DRB1" > hla_regions.bed
# Get primary contigs as well as alternative regions for the current reference
cut -f 1,2 ../reference/GRCh38_full_analysis_set_plus_decoy_hla.fa.fai | grep -w -E "^chr([1-9]|1[0-9]|2[0-2]|[XYM])" | awk '{print $1,0,$2}' OFS="\t" > no_alts.bed
cut -f 1,2 ../reference/GRCh38_full_analysis_set_plus_decoy_hla.fa.fai | grep -w -v -E "^chr([1-9]|1[0-9]|2[0-2]|[XYM])" | awk '{print $1,0,$2}' OFS="\t" > alts.bed
# Extract sequences
bedtools getfasta -fi ../reference/GRCh38_full_analysis_set_plus_decoy_hla.fa -bed no_alts.bed | sed 's/\(.*\):[^:]*$/\1/' > GRCh38_no_alts.fa
bedtools getfasta -fi ../reference/GRCh38_full_analysis_set_plus_decoy_hla.fa -bed alts.bed | sed 's/\(.*\):[^:]*$/\1/' > GRCh38_alts.fa
# Align alternative contigs to primary chromosomes
minimap2 -t 8 -x asm5 --eqx -c GRCh38_no_alts.fa GRCh38_alts.fa | bgzip -c > GRCh38_alts_to_no_alts.paf.gz
bgzip -r GRCh38_alts_to_no_alts.paf.gz
# Query target regions
impg query -p GRCh38_alts_to_no_alts.paf.gz -b hla_regions.bed > hla_regions.bedpe
# Merge primary and alternative regions
awk '
NR==FNR {
region = $1":"$2"-"$3
simple_region = $1 "\t" $2 "\t" $3
regions[region] = $0
regions_simple[simple_region] = $0
labels[$1 "\t" $2 "\t" $3] = (NF >= 4) ? $4 : ""
next
}
{
# Try to match full coordinate format first (chr6:start-end)
query_region = $7
allele = $1":"$2"-"$3
# If column 7 is not in coordinate format, construct it from columns 4-6
if (query_region !~ /:/) {
query_region = $4":"$5"-"$6
}
if (query_region in regions) {
if (alleles[query_region]) {
alleles[query_region] = alleles[query_region] "," allele
} else {
alleles[query_region] = allele
}
}
}
END {
for (region in regions) {
split(regions[region], r, "\t")
label = (r[4] != "") ? r[4] : ""
if (label != "") {
print r[1], r[2], r[3], label, alleles[region]
} else {
print r[1], r[2], r[3], alleles[region]
}
}
}
' OFS="\t" hla_regions.bed hla_regions.bedpe | bedtools sort -i > hla_roi.bed
cd ..
# Annotations (optional)
You can optionally provide gene annotation and protein translation files to the pipeline. If supplied, the pipeline will build gene graphs using pangene (opens new window) allowing gene projection onto the assemblies for the regions of interest.
mkdir annotations
cd annotations
# Download GTF annotation
wget https://ftp.ebi.ac.uk/pub/databases/gencode/Gencode_human/release_48/gencode.v48.annotation.gtf.gz
# Download and index protein-coding transcript translations
wget https://ftp.ebi.ac.uk/pub/databases/gencode/Gencode_human/release_48/gencode.v48.pc_translations.fa.gz
gzip -d gencode.v48.pc_translations.fa.gz
bgzip gencode.v48.pc_translations.fa
samtools faidx gencode.v48.pc_translations.fa.gz
cd ..
# Haplotype exclusion (optional)
You may optionally provide a list of regions identified by flagger (opens new window) in BED format to exclude potentially misassembled haplotype blocks from downstream analyses. The following strategy downloads the Flagger index and corresponding BED files for our assemblies of interest, then generates a unified exclusion file:
mkdir flagger
cd flagger
# Download Flagger index
wget https://raw.githubusercontent.com/human-pangenomics/HPP_Year1_Assemblies/12656b82a42cd4ec6d421abe7fd4ebdeca74b41c/annotation_index/Year1_assemblies_v2_genbank_Flagger.index
# Extract assembly names and download corresponding BEDs
cut -f 1 ../assemblies_a/*fai | cut -d "#" -f 1 | sort | uniq | while read f; do
link=$(grep -w $f Year1_assemblies_v2_genbank_Flagger.index | cut -f 2)
clink=$(echo $link | sed 's,s3://,https://s3-us-west-2.amazonaws.com/,')
wget $clink
done
# Merge and sort all exclusion regions
cut -f 1-3 *bed | bedtools sort -i - > flagger.exclude.bed
cd ..
# Clone cosigt Repository
Clone the cosigt repository from GitHub (opens new window).
micromamba activate smk7324app132
git clone https://github.com/davidebolo1993/cosigt
cd cosigt
For the environment setup, refer to the → setup section.
# Configure and run the workflow (master and ancient_dna branches)
With all necessary data prepared, we can now configure the pipeline using the dedicated setup script (opens new window). The organize.py script of the master and ancient_dna branches of the pipeline accepts the arguments listed below:
# Confirm you're on the master branch
git rev-parse --abbrev-ref HEAD
# Should return 'master'
cd cosigt_smk
python workflow/scripts/organize.py -h
COsine SImilarity-based GenoTyper
optional arguments:
-h, --help show this help message and exit
Required I/O arguments:
-a FASTA, --assemblies FASTA
assemblies individuals to -r will be genotyped against. This is a tab-separated file mapping chromosomes in -b to a FASTA with contigs for that chromosome. FASTA can be bgzip-compressed and must be indexed
-g FASTA, --genome FASTA
reference genome. This is the FASTA regions to -b refers to. FASTA can be bgzip-compressed and must be indexed
-r TSV, --reads TSV individuals to genotype. This is a tab-separated file mapping each alignment file (1st column, BAM/CRAM, indexed) to an id (2nd column)
-b BED, --bed BED regions to genotype. A standard 3-column BED file, but can have a 4th column to label the region and a 5th column listing comma-separated alternative contigs for that region
-o FOLDER, --output FOLDER
output folder. This is where results from cosigt pipeline will be stored
Optional arguments:
--gtf Gene annotation on the reference chromosomes in GTF format [None]
--proteins Protein-coding transcript translation sequences in FASTA format. FASTA can be bgzip-compressed and must be indexed [None]
--tmp tmp directory. Will be used by singularity and pggb [/tmp]
--pansn PanSN prefix naming for the reference genome [grch38#1#]
--profile snakemake profile, if available [None]
--flagger regions to exclude for each contigs. This is a standard BED file coming from flagger, with contigs names matching those of the assemblies in -a [None]
--conda prepare for running using conda instead of singularity [False]
--threads run snakemake using that many cores - ignored if using a profile [32]
--no_pangene DO NOT visualize coding genes on input haplotypes [False]
--no_odgi DO NOT visualize node coverage on input haplotypes in a "odgi viz"-like format [False]
--wally run wally to visualize reads-to-haplotypes realignment [False]
--svbyeye run SVByEye to visualize predicted haplotypes-to-reference realignment [False]
For the current use-case, inputs to organize.py are the following:
# Assemblies map
> ../../asm_map.tsv
for f in ../../assemblies_a/*.fasta.gz; do
id=$(basename "$f" | cut -d "." -f 1)
echo -e "$id\t$f" >> ../../asm_map.tsv
done
assemblies=$(readlink -f ../../asm_map.tsv)
# Reference genome
genome=$(readlink -f ../../reference/GRCh38_full_analysis_set_plus_decoy_hla.fa)
# Read alignments
> ../../aln_map.tsv
for r in ../../alignments/*cram; do
id=$(basename "$r" | cut -d "." -f 1)
echo -e "$r\t$id" >> ../../aln_map.tsv
done
reads=$(readlink -f ../../aln_map.tsv)
# Genotyping regions
bed=$(readlink -f ../../regions/roi.bed)
# Output directory
output=$(readlink -f $PWD/cosigt_results)
# GTF (optional)
gtf=$(readlink -f ../../annotations/gencode.v48.annotation.gtf.gz)
# Proteins FASTA (optional)
proteins=$(readlink -f ../../annotations/gencode.v48.pc_translations.fa.gz)
# Flagger exclusion (optional)
flagger=$(readlink -f ../../flagger/flagger.exclude.bed)
We can then run the script:
# Clean resources
if [ -d resources ]; then
rm -rf resources
fi
# Run organize.py
python workflow/scripts/organize.py \
-a $assemblies \
-g $genome \
-r $reads \
-b $bed \
-o $output \
--gtf $gtf \
--proteins $proteins \
--flagger $flagger \
--wally \
--svbyeye \
--threads 8
If a profile was generated as explained in the → setup section, one can provide such profile to the --profile flag (for instance, --profile config/slurm).
This generates in the current working directory (cosigt/cosigt_smk):
- the
resourcesfolder - the
config/config.yamlfile - the
cosigt_smk.shscript
In resources input files are re-organized so that the pipeline knows exactly where they are located. They follow the structure below:
tree resources
resources/
|-- alignments
| |-- HG00438.cram -> /<path>/cosigt_test/alignments/HG00438.final.cram
| |-- HG00438.cram.crai -> /<path>/cosigt_test/alignments/HG00438.final.cram.crai
| |-- HG01952.cram -> /<path>/cosigt_test/alignments/HG01952.final.cram
| `-- HG01952.cram.crai -> /<path>/cosigt_test/alignments/HG01952.final.cram.crai
|-- annotations
| |-- gencode.v48.annotation.gtf.gz -> /<path>/cosigt_test/annotations/gencode.v48.annotation.gtf.gz
| |-- gencode.v48.pc_translations.fa.gz -> /<path>/cosigt_test/annotations/gencode.v48.pc_translations.fa.gz
| |-- gencode.v48.pc_translations.fa.gz.fai -> /<path>/cosigt_test/annotations/gencode.v48.pc_translations.fa.gz.fai
| `-- gencode.v48.pc_translations.fa.gz.gzi -> /<path>/cosigt_test/annotations/gencode.v48.pc_translations.fa.gz.gzi
|-- assemblies
| |-- chr4
| | |-- chr4.subset.fasta.gz -> /<path>/cosigt_test/assemblies_a/chr4.subset.fasta.gz
| | |-- chr4.subset.fasta.gz.fai -> /<path>/cosigt_test/assemblies_a/chr4.subset.fasta.gz.fai
| | `-- chr4.subset.fasta.gz.gzi -> /<path>/cosigt_test/assemblies_a/chr4.subset.fasta.gz.gzi
| `-- chr6
| |-- chr6.subset.fasta.gz -> /<path>/cosigt_test/assemblies_a/chr6.subset.fasta.gz
| |-- chr6.subset.fasta.gz.fai -> /<path>/cosigt_test/assemblies_a/chr6.subset.fasta.gz.fai
| `-- chr6.subset.fasta.gz.gzi -> /<path>/cosigt_test/assemblies_a/chr6.subset.fasta.gz.gzi
|-- flagger
| `-- flagger_blacklist.bed
|-- reference
| |-- GRCh38_full_analysis_set_plus_decoy_hla.fa -> /<path>/cosigt_test/reference/GRCh38_full_analysis_set_plus_decoy_hla.fa
| `-- GRCh38_full_analysis_set_plus_decoy_hla.fa.fai -> /<path>/cosigt_test/reference/GRCh38_full_analysis_set_plus_decoy_hla.fa.fai
`-- regions
|-- all_regions.tsv
|-- chr4
| `-- chr4_144109303_144140751.bed
`-- chr6
`-- chr6_31972057_32055418.bed
The config/config.yaml file defines resources for pipeline components. For example:
grep -A 5 "pggb" config/config.yaml
pggb:
mem_mb: 20000
params: -c 2 -k 101
threads: 24
time: 40
tmpdir: /tmp
This configures pggb to run with 20GB RAM, 24 threads, and 40 minutes time limit using /tmp as a working directory (/tmp can be changed by passing a different directory to --tmp argument of the organize.py script). Resources can be tuned by users based on their availability.
The cosigt_smk.sh script launches the pipeline. By default, the pipeline will be launched using Singularity for tools. Since we haven’t passed a --profile, the workflow will execute locally. To run on a SLURM cluster, pass the profile path (see the → setup section for details on how to build an example profile for SLURM).
Note that users may want to refine the regions provided to --bed using the refine mode of cosigt. This is not required for the current example.
Cosigt pipeline can be run with:
sh cosigt_smk.sh
# Exploring Results
Following, an overview of the primary outputs produced by the cosigt pipeline.
# Genotyping Results
Cosigt performs haplotype deconvolution, that is identifies the combination of haplotypes from the pangenome that best matches the short-read sequencing data for a given region. Details on this step are described in the → workflow section. After running the pipeline, genotyping outputs are stored under:
tree cosigt_results/cosigt
cosigt_results/cosigt/
|-- HG00438
| |-- chr4
| | `-- chr4_144109303_144140751
| | |-- chr4_144109303_144140751.cosigt_genotype.tsv
| | |-- chr4_144109303_144140751.sorted_combos.tsv.gz
| | `-- viz
| | |-- chr4_144109303_144140751.ava.png
| | |-- HG00438#1_to_grch38#1#chr4.png
| | `-- HG00438#2_to_grch38#1#chr4.png
| `-- chr6
| `-- chr6_31972057_32055418
| |-- chr6_31972057_32055418.cosigt_genotype.tsv
| |-- chr6_31972057_32055418.sorted_combos.tsv.gz
| `-- viz
| |-- chr6_31972057_32055418.ava.png
| |-- HG00438#1_to_grch38#1#chr6.png
| `-- HG00438#2_to_grch38#1#chr6.png
`-- HG01952
|-- chr4
| `-- chr4_144109303_144140751
| |-- chr4_144109303_144140751.cosigt_genotype.tsv
| |-- chr4_144109303_144140751.sorted_combos.tsv.gz
| `-- ...
`-- chr6
`-- chr6_31972057_32055418
|-- chr6_31972057_32055418.cosigt_genotype.tsv
|-- chr6_31972057_32055418.sorted_combos.tsv.gz
`-- ...
Each region directory contains:
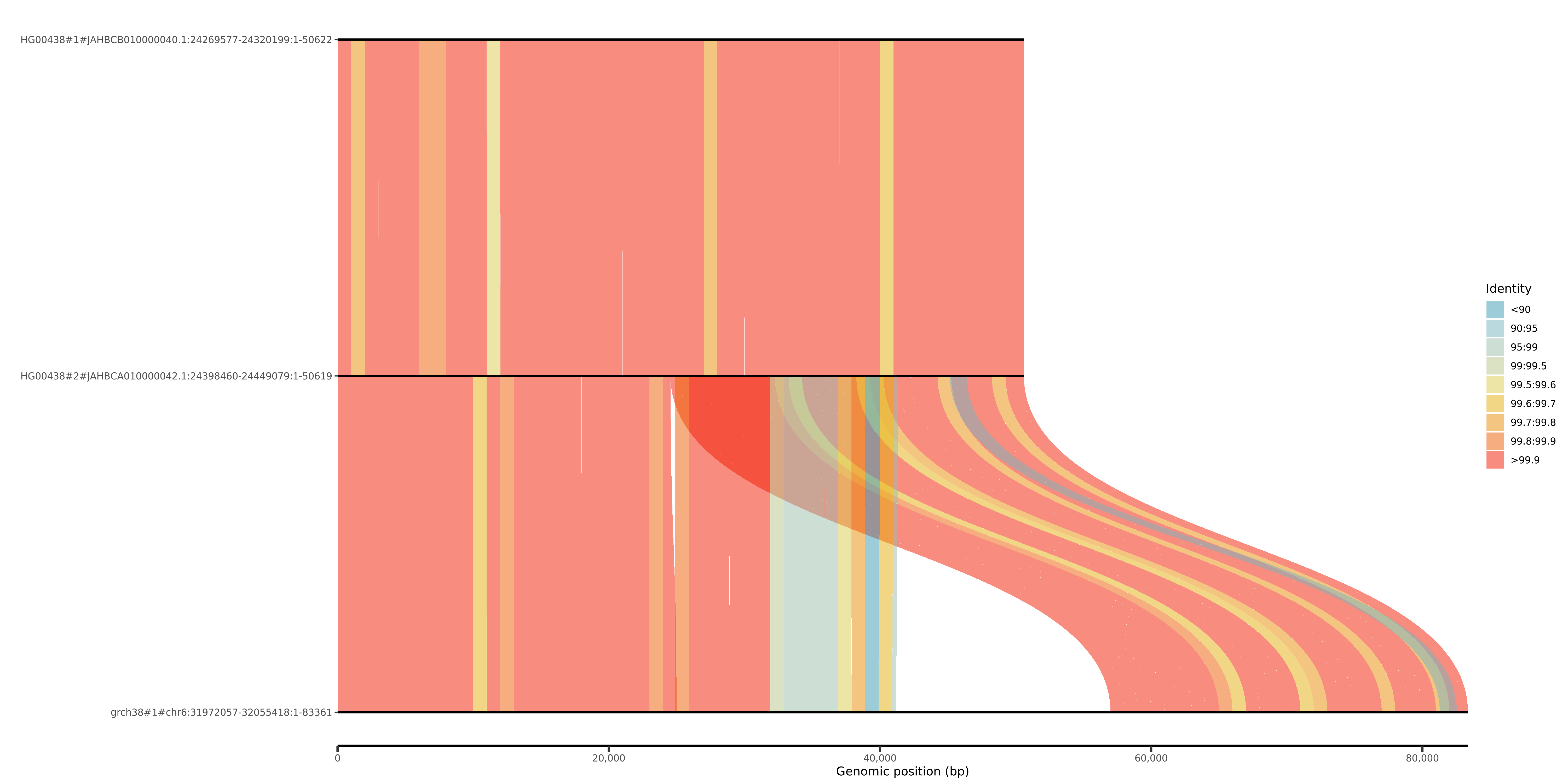
*.cosigt_genotype.tsv: best-matching haplotype pair (haplotype.1,haplotype.2) selected to maximize the cosine similarity (opens new window) to the sample.*.sorted_combos.tsv.gz: cosine similarity scores for all tested haplotype pairs.viz/: visualizations of alignments between predicted haplotypes and the reference (only if--svbyeyeis provided toorganize.py) One example forHG00438at the C4 locus (chr6_31972057_32055418):
In this case, both predicted haplotypes (HG00438#1, HG00438#2) are structurally similar to each other but diverge from the reference (grch38#chr6), due to a duplication absent in the predicted haplotypes.
# Structural Clustering
To summarize major structural patterns, cosigt performs clustering of haplotypes based on graph similarity (see the → workflow section). Outputs are stored in:
tree cosigt_results/cluster
cosigt_results/cluster/
|-- chr4
| `-- chr4_144109303_144140751
| |-- chr4_144109303_144140751.clusters.hapdist.norm.tsv
| |-- chr4_144109303_144140751.clusters.hapdist.tsv
| |-- chr4_144109303_144140751.clusters.json
| |-- chr4_144109303_144140751.clusters.medoids.tsv
| |-- chr4_144109303_144140751.clusters.metrics.tsv
| `-- chr4_144109303_144140751.clusters.tsv
`-- chr6
`-- chr6_31972057_32055418
|-- chr6_31972057_32055418.clusters.hapdist.norm.tsv
|-- chr6_31972057_32055418.clusters.hapdist.tsv
|-- chr6_31972057_32055418.clusters.json
|-- chr6_31972057_32055418.clusters.medoids.tsv
|-- chr6_31972057_32055418.clusters.metrics.tsv
`-- chr6_31972057_32055418.clusters.tsv
File descriptions:
*.clusters.tsv/*.clusters.json: assignedhaplotype.groupperhaplotype.name(TSV and JSON formats).*.clusters.hapdist.tsv: pairwise haplotype distances between groups.*.clusters.hapdist.norm.tsv: normalized version of the above (scaled to max pairwise distance).*.clusters.medoids.tsv: representative haplotype (medoid) for each group.*.clusters.metrics.tsv: clustering parameters (e.g., eps) used.
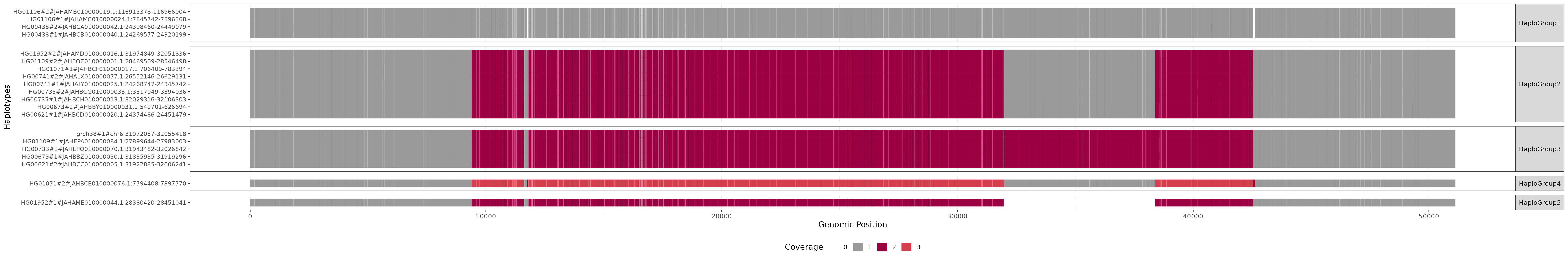
Visualizations of clusters can be found in cosigt_test/odgi/viz/{chromosome}/{region}. An example for the C4 locus:
Nodes are oriented relative to the reference (grch38#1#chr6) and colored by path coverage (based on the Spectral-11 (opens new window) palette). Haplotypes are grouped and ordered by cluster, facilitating visual comparison of structural similarity.
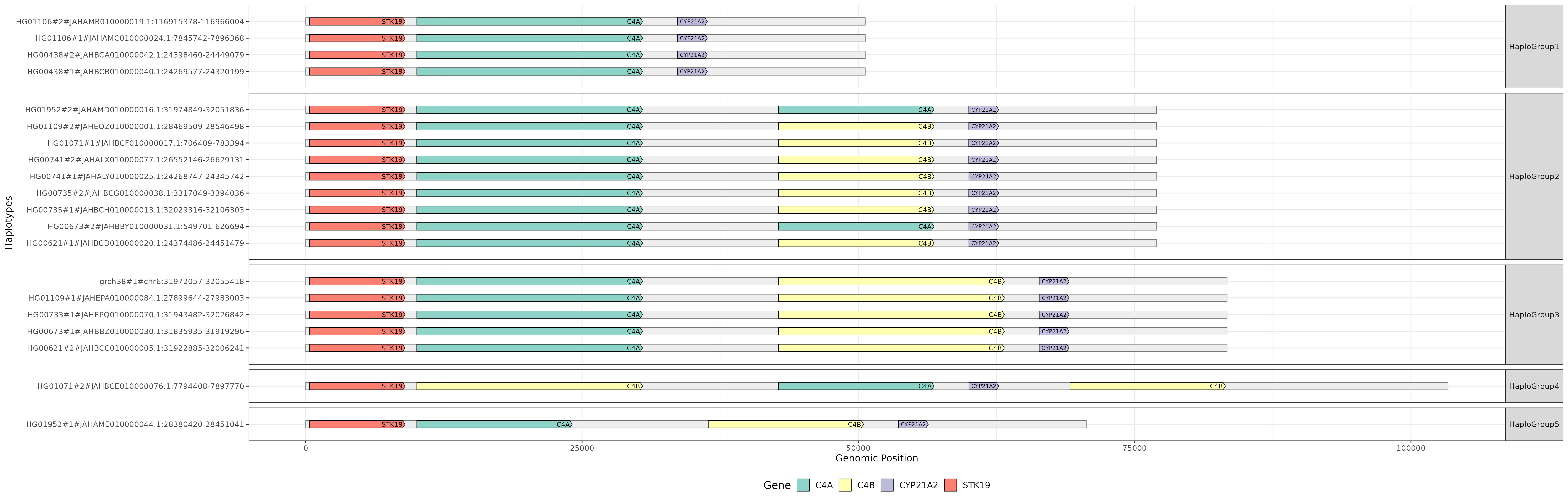
# Genes annotation
If gene annotations and protein sequences were provided, cosigt uses pangene to build gene graphs and project annotations onto the assemblies. An example visualization for the C4 locus:
# Read Realignment Visualization
When the --wally flag is provided to organize.py, cosigt generates read realignment visualizations for each assembly block. These plots display how individual reads align to the assembled haplotype blocks. The visualization files are organized in the following directory structure:
tree cosigt_results/wally/HG00438/chr6/chr6_31972057_32055418
cosigt_results/wally/HG00438/chr6/chr6_31972057_32055418
|-- grch38#1#chr6:31972057-32055418_1_83361.png
|-- HG00438#1#JAHBCB010000040.1:24269577-24320199_1_50622.png
|-- HG00438#2#JAHBCA010000042.1:24398460-24449079_1_50619.png
`-- ...
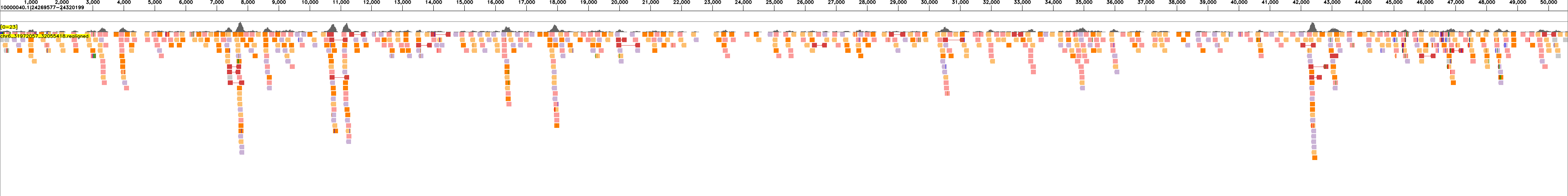
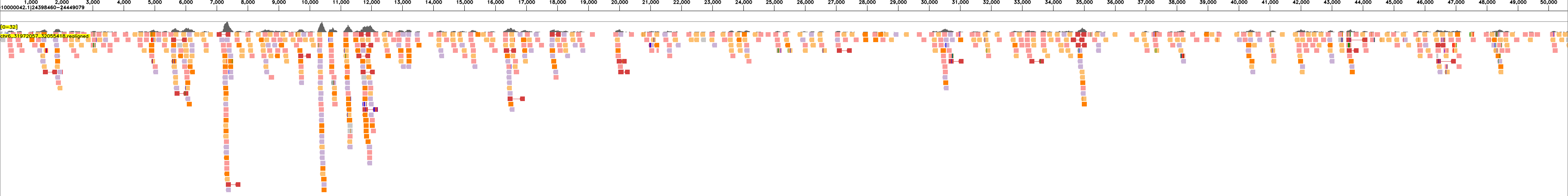
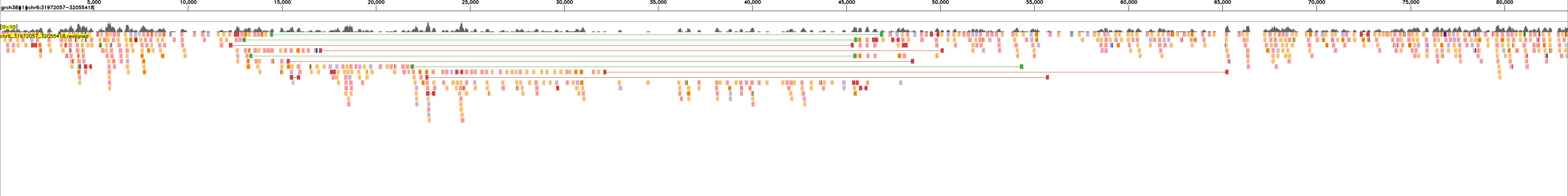
The following examples show read realignments for sample HG00438 to the expected haplotype blocks:
and to the reference:
These visualizations enable quality assessment of the assemblies by showing the concordance between sequencing reads and the assembled haplotype blocks.
# Interactive plot of genotyping results
The cosigt pipeline includes an interactive visualization tool (opens new window) that enables dynamic exploration of genotyping outcomes across samples and regions. You can launch the app as follows:
cd workflow/scripts
bash runapp.sh ../../cosigt_test
#Listening on http://0.0.0.0:3838
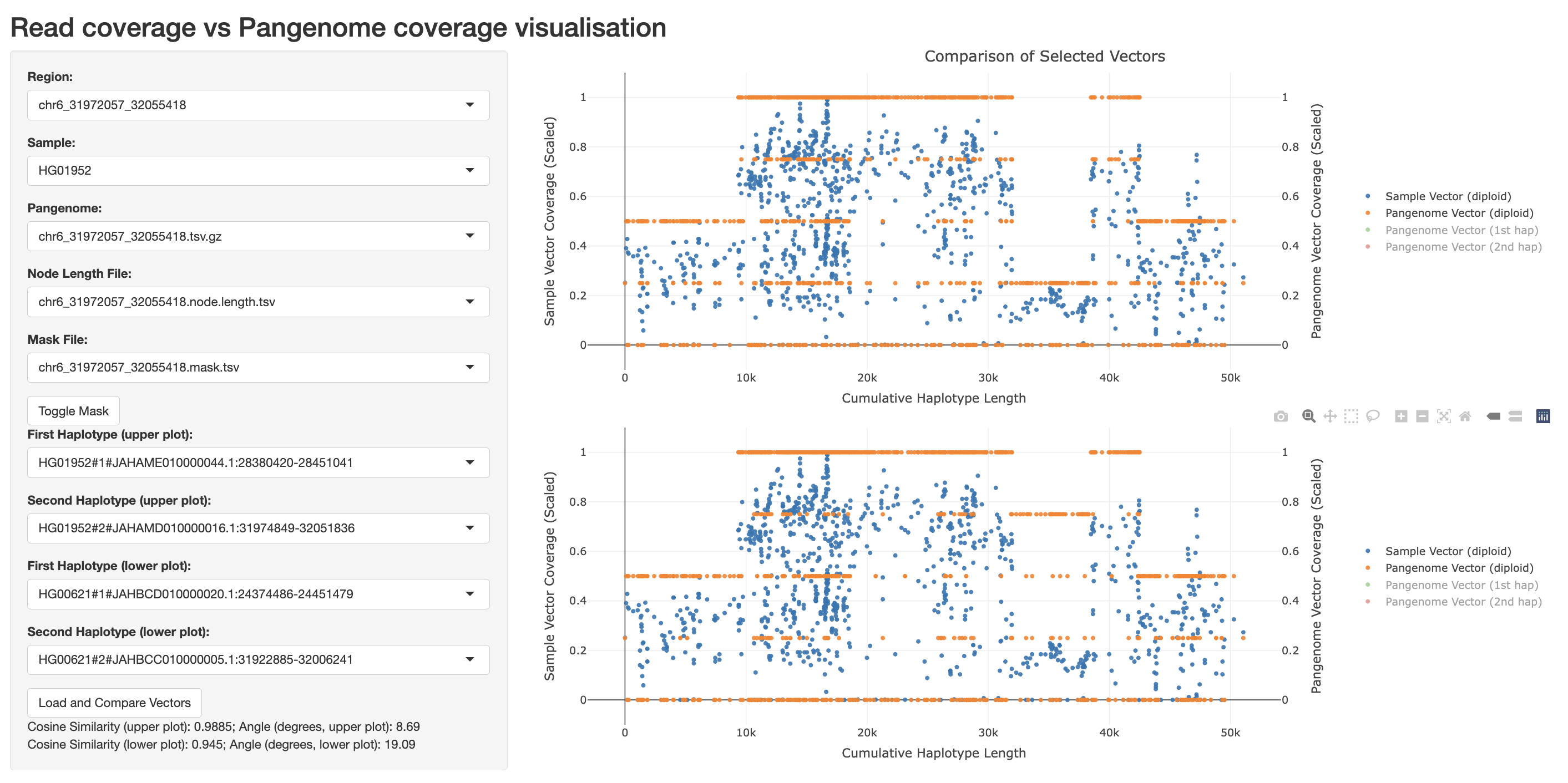
Then, open your web browser and navigate to http://0.0.0.0:3838 to view the results. For example:
The app primarily visualizes short-read coverage (Sample Vector Coverage, blue dots, left y-axis) versus haplotype coverage (Pangenome Vector Coverage, orange dots, right y-axis) across graph nodes (Cumulative Haplotype Length, x-axis). Each dot represents a graph node, and the Cumulative Haplotype Length accounts for the length of each node.
For each region of interest (selectable via the Region menu) and sample (via the Sample menu), you can display up to two short-read vs haplotype coverage comparisons: the first in the upper panel and the second in the lower panel. This allows you to compare the cosigt genotyping prediction against the expected haplotype coverage, helping to identify matches or coverage anomalies.
# Configure and run the workflow (custom_alleles branch)
As an alternative to running from raw assemblies, cosigt can also be launched directly from a predefined set of alleles — for example, those generated using different methods (that is, not via impg) or in a previous run:
tree cosigt_results/bedtools/getfasta
cosigt_results/bedtools/getfasta/
|-- chr4
| `-- chr4_144109303_144140751
| |-- chr4_144109303_144140751.fasta.gz
| |-- chr4_144109303_144140751.fasta.gz.fai
| `-- chr4_144109303_144140751.fasta.gz.gzi
`-- chr6
`-- chr6_31972057_32055418
|-- chr6_31972057_32055418.fasta.gz
|-- chr6_31972057_32055418.fasta.gz.fai
`-- chr6_31972057_32055418.fasta.gz.gzi
To use this mode, switch to the custom_alleles branch:
cd ../
git checkout custom_alleles
cd cosigt_smk
python workflow/scripts/organize.py -h
The script accepts a similar set of arguments to the master and ancient_dna branches. The key difference lies in the -a argument, which now expects a TSV mapping each region to its corresponding allele FASTA file (can be bgzipped). Example:
> ../../all_map.tsv
for f in cosigt_results/bedtools/getfasta/*/*/*gz; do
id=$(basename $(dirname $f))
echo -e "$id\t$f" >> ../../all_map.tsv
done
Then run:
# Clean previous resources
rm -rf resources
# Alleles
alleles=$(readlink -f ../../all_map.tsv)
# New output directory
output=$(readlink -f $PWD/cosigt_results_alleles)
# Launch organize.py
python workflow/scripts/organize.py \
-a $alleles \
-g $genome \
-r $reads \
-b $bed \
-o $output \
--gtf $gtf \
--proteins $proteins \
--flagger $flagger \
--threads 8
And finally:
sh cosigt_smk.sh
The output structure and interpretation remain identical to those described in the previous sections.
# Refine Workflow
In addition to the main cosigt workflow, our Snakemake pipeline also supports a refine mode (available in both the master and ancient_dna branches).
This mode attempts to dynamically extend the flanks of input regions to retain as many haplotypes as possible during the impg step.
To run the refine workflow, simply replace "cosigt" with "refine" in the cosigt_smk.sh.
# Clean resources
rm -rf resources
# Output directory
output=$(readlink -f $PWD/cosigt_results_refine)
# Run organize.py
python workflow/scripts/organize.py \
-a $assemblies \
-g $genome \
-r $reads \
-b $bed \
-o $output
#change cosigt to refine
sed -i 's/ cosigt --/ refine --/' cosigt_smk.sh
#run
sh cosigt_smk.sh
The output folder should look like this:
tree cosigt_results_refine
cosigt_results_refine
|-- impg
| |-- chr4
| | `-- chr4.paf.gz.impg
| `-- chr6
| `-- chr6.paf.gz.impg
|-- minimap2
| |-- chr4
| | |-- chr4.fasta.gz
| | |-- chr4.fasta.gz.fai
| | |-- chr4.fasta.gz.gzi
| | |-- chr4.paf.gz
| | `-- chr4.paf.gz.gzi
| `-- chr6
| |-- chr6.fasta.gz
| |-- chr6.fasta.gz.fai
| |-- chr6.fasta.gz.gzi
| |-- chr6.paf.gz
| `-- chr6.paf.gz.gzi
`-- refine
|-- impg
| |-- chr4
| | `-- chr4_144109303_144140751
| | |-- chr4_144109303_144140751.haplotypes.bed
| | `-- chr4_144109303_144140751.refined.bed
| `-- chr6
| `-- chr6_31972057_32055418
| |-- chr6_31972057_32055418.haplotypes.bed
| `-- chr6_31972057_32055418.refined.bed
`-- regions_refined.bed
The main output is a BED file containing the refined regions in cosigt_results_refine/refine/regions_refined.bed. This refined BED file can then be used as input for a new cosigt run by replacing the original regions BED file - even if in this specific example there is no difference between the original regions and those processed by the refine module (as shown with diff <(cat cosigt_results_refine/refine/regions_refined.bed) <(cat resources/regions/all_regions.tsv)):
# Clean resources
rm -rf resources
# Output directory
output=$(readlink -f $PWD/cosigt_results_refine)
bed=$(readlink -f $PWD/cosigt_results_refine/refine/regions_refined.bed)
# Run organize.py
python workflow/scripts/organize.py \
-a $assemblies \
-g $genome \
-r $reads \
-b $bed \
-o $output
#run
sh cosigt_smk.sh
By re-using the same output directory, the pipeline will skip the initial query-to-target alignment step, recycling those generated in the refine mode.